(New page: [http://balthier.ecn.purdue.edu/index.php/ECE662 ECE662 Main Page] [http://balthier.ecn.purdue.edu/index.php/ECE662#Class_Lecture_Notes Class Lecture Notes] Nearest Neighbors Clarificati...) |
|||
| (41 intermediate revisions by 8 users not shown) | |||
| Line 1: | Line 1: | ||
| − | [ | + | =Lecture 18, [[ECE662]]: Decision Theory= |
| + | Lecture notes for [[ECE662:BoutinSpring08_Old_Kiwi|ECE662 Spring 2008]], Prof. [[user:mboutin|Boutin]]. | ||
| − | + | Other lectures: [[Lecture 1 - Introduction_Old Kiwi|1]], | |
| + | [[Lecture 2 - Decision Hypersurfaces_Old Kiwi|2]], | ||
| + | [[Lecture 3 - Bayes classification_Old Kiwi|3]], | ||
| + | [[Lecture 4 - Bayes Classification_Old Kiwi|4]], | ||
| + | [[Lecture 5 - Discriminant Functions_Old Kiwi|5]], | ||
| + | [[Lecture 6 - Discriminant Functions_Old Kiwi|6]], | ||
| + | [[Lecture 7 - MLE and BPE_Old Kiwi|7]], | ||
| + | [[Lecture 8 - MLE, BPE and Linear Discriminant Functions_Old Kiwi|8]], | ||
| + | [[Lecture 9 - Linear Discriminant Functions_Old Kiwi|9]], | ||
| + | [[Lecture 10 - Batch Perceptron and Fisher Linear Discriminant_Old Kiwi|10]], | ||
| + | [[Lecture 11 - Fischer's Linear Discriminant again_Old Kiwi|11]], | ||
| + | [[Lecture 12 - Support Vector Machine and Quadratic Optimization Problem_Old Kiwi|12]], | ||
| + | [[Lecture 13 - Kernel function for SVMs and ANNs introduction_Old Kiwi|13]], | ||
| + | [[Lecture 14 - ANNs, Non-parametric Density Estimation (Parzen Window)_Old Kiwi|14]], | ||
| + | [[Lecture 15 - Parzen Window Method_Old Kiwi|15]], | ||
| + | [[Lecture 16 - Parzen Window Method and K-nearest Neighbor Density Estimate_Old Kiwi|16]], | ||
| + | [[Lecture 17 - Nearest Neighbors Clarification Rule and Metrics_Old Kiwi|17]], | ||
| + | [[Lecture 18 - Nearest Neighbors Clarification Rule and Metrics(Continued)_Old Kiwi|18]], | ||
| + | [[Lecture 19 - Nearest Neighbor Error Rates_Old Kiwi|19]], | ||
| + | [[Lecture 20 - Density Estimation using Series Expansion and Decision Trees_Old Kiwi|20]], | ||
| + | [[Lecture 21 - Decision Trees(Continued)_Old Kiwi|21]], | ||
| + | [[Lecture 22 - Decision Trees and Clustering_Old Kiwi|22]], | ||
| + | [[Lecture 23 - Spanning Trees_Old Kiwi|23]], | ||
| + | [[Lecture 24 - Clustering and Hierarchical Clustering_Old Kiwi|24]], | ||
| + | [[Lecture 25 - Clustering Algorithms_Old Kiwi|25]], | ||
| + | [[Lecture 26 - Statistical Clustering Methods_Old Kiwi|26]], | ||
| + | [[Lecture 27 - Clustering by finding valleys of densities_Old Kiwi|27]], | ||
| + | [[Lecture 28 - Final lecture_Old Kiwi|28]], | ||
| + | ---- | ||
| + | ---- | ||
| − | Nearest Neighbors | + | == Nearest Neighbors Classification Rule (Alternative Approach) == |
| − | --[[ | + | |
| + | * Find invariant coordinates | ||
| + | <math>\varphi : \Re ^k \rightarrow \Re ^n </math> | ||
| + | such that <math>\varphi (x) = \varphi (\bar x) </math> for all <math>x, \bar x</math> which are related by a rotation & translation | ||
| + | Do NOT trivialize! | ||
| + | |||
| + | '''Example:''' <math>\varphi (x) =0 </math> gives us a trivial invariant coordinate. But, you lose information about separation, since everything is mapped to zero. | ||
| + | |||
| + | Want <math>\varphi (x) = \varphi (\bar x) </math> | ||
| + | <math>\Leftrightarrow x, \bar x</math> are related by a rotation and translation | ||
| + | |||
| + | '''Example:''' | ||
| + | <math>p=(p_1,p_2,\cdots, p_N) \in \Re ^{3 \times N}</math> | ||
| + | <math>\varphi</math> maps representation position of tags on body onto <math>(d_{12},d_{13},d_{14},\cdots , d_{N-1, N} )</math> | ||
| + | where <math>d_{ij}</math>= Euclidean distance between <math>p_i</math> and <math>p_j</math> | ||
| + | |||
| + | In the above example, we can reconstruct up to a rotation and translation. | ||
| + | |||
| + | WARNING: Euclidean distance in the invariant coordinate space has nothing to do with Euclidean distance or Procrustes distance in initial feature space. | ||
| + | |||
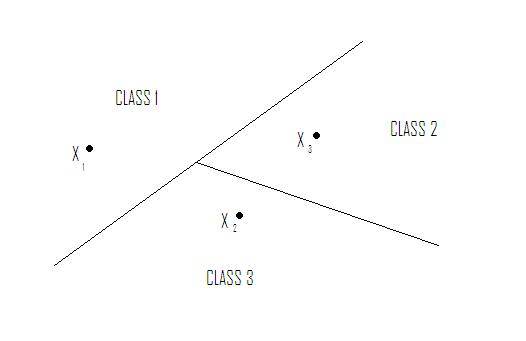
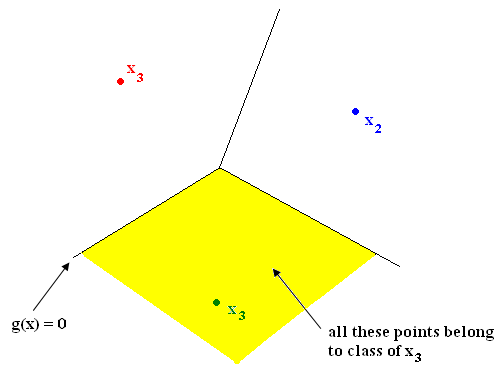
| + | Nearest Neighbor in <math>\Re ^2</math> yields tessellation (tiling of floor with 2D shapes such that 1) no holes and 2) cover all of <math>\Re ^2</math> ). The tessellations separate sample space into regions. Shape of cells depends on metric chosen. See Figure 1. | ||
| + | |||
| + | |||
| + | <center>[[Image:Tessellation_Old Kiwi.jpg|frame|Figure 1 - Separation of Sample Space using Tessellations]]</center> | ||
| + | |||
| + | |||
| + | <center>[[Image:lec18_nn_r2_Old Kiwi.PNG|frame|Figure 1b - Tessellations]]</center> | ||
| + | |||
| + | |||
| + | '''Example:''' if feature vectors are such that vectors related by a rotation belong to same class | ||
| + | <math>\rightarrow</math> metric should be chosen so that tiles are rotationally symmetric. See Figure 2. | ||
| + | |||
| + | |||
| + | <center>[[Image:Rotation_Old Kiwi.jpg|frame|Figure 2 - Example of Vectors related by Rotations]]</center> | ||
| + | |||
| + | |||
| + | Instead of working with (x,y) rotationally invariant, work with <math>z=\sqrt{x^2 + y^2}</math> (distance from origin) | ||
| + | |||
| + | How good is Nearest Neighbor rule? | ||
| + | # Training error is zero: does not measure the "goodness" of a rule | ||
| + | # Test error: want it to be equal to Bayes error rate, because this yields the minimum error | ||
| + | |||
| + | ---- | ||
| + | |||
| + | == Nearest Neighbor error rate == | ||
| + | |||
| + | '''Recall:''' | ||
| + | Probability of error (error rate) on test data is | ||
| + | <math>P(e)=\int p(e \mid \vec{x}) p(\vec{x}) d\vec{x}</math> | ||
| + | |||
| + | Let <math>P_d(e)</math> be the error rate when d training samples are used. | ||
| + | |||
| + | Let <math>P=\lim_{d \rightarrow \infty } P_{d}(e) </math> | ||
| + | |||
| + | '''Claim:''' limit error rate <math>P=\int (1-\sum _{i=1} ^{c}p^2 (\omega _i \mid \vec{x}))p(x)dx</math> | ||
| + | |||
| + | '''Proof of claim:''' Given observation <math>\vec{x}</math>, denode by <math>\vec{x'}_d</math> the nearest neighbor of <math>\vec{x}</math> among <math>\{\vec{x}_1,\vec{x}_2, \cdots , \vec{x}_d \}</math> | ||
| + | |||
| + | <math>P_d(e \mid \vec{x})=\int p_d(e \mid \vec{x}, \vec{x'}_d)p_d(\vec{x'}_d \mid \vec{x})p(x)dx</math> | ||
| + | but <math>\lim _{d \rightarrow \infty } p_d (\vec{x'}_d \mid \vec{x})=\delta ({\vec{x' _d }-\vec{x}})</math> | ||
| + | |||
| + | because probability that sample falls into region R centered at <math>\vec{x}</math> is <math>P_{R}=\int _R p(\vec{x' _d})d \vec{x'}_d</math>. | ||
| + | |||
| + | So, if <math>p(\vec{x}) \neq 0</math> (true almost everywhere), then probability that all samples fall outside R is <math>\lim _{d \rightarrow \infty} | ||
| + | {(1-P_{R})}^d =0</math> | ||
| + | |||
| + | So, <math>\lim _{d \rightarrow \infty} \vec{x'}_d = \vec{x}</math> and <math>p_d (\vec{x'}_d \mid \vec{x})=\delta (\vec{x'}_d -\vec{x})+\epsilon _d (\vec{x})</math> where <math>\lim _{d \rightarrow \infty } \epsilon _d (x)=0</math> | ||
| + | |||
| + | Now <math>p_d (e \mid \vec{x}, \vec{x'}_d)</math> = ? | ||
| + | |||
| + | Let <math>\theta , \theta _1 , \theta _2 , \cdots, \theta _{d}</math> be the class of <math>x , x_1 , x_2 , \cdots , x_d</math>, respectively. | ||
| + | |||
| + | Using nearest neighbor rule, error if <math>\theta \neq</math>class of <math> \vec{x'}_d</math><math>=: \theta ' _d</math> | ||
| + | |||
| + | <math>\Rightarrow p_d(e \mid \vec{x},\vec{x'}_d)=1-\sum_{i=1} ^ c p(\theta = \omega _i , \theta ' _d = \omega _i \mid \vec{x}, \vec{x'}_d )</math> | ||
| + | |||
| + | <math>=1- \sum _{i=1} ^c p(\omega _i \mid \vec{x}) p(\omega _i \mid \vec{x'} _d) </math> | ||
| + | |||
| + | Recall <math>p_d (e \mid \vec{x}, \vec{x'}_d)p_d (\vec{x'}_d \mid \vec{x})d \vec{x'}_d</math> | ||
| + | |||
| + | You get: <math>p_d (e \mid \vec{x})=(1-\sum _{i=1} ^c p(\omega _i \mid x) p(\omega _i \mid x) )</math> + {something that goes to zero as d goes to <math>\infty</math>} | ||
| + | |||
| + | <math>\lim _{d \rightarrow \infty} p_d (e \mid \vec{x})=(1- \sum _{i=1} ^c {p(\omega _i \mid x)}^2)</math> | ||
| + | ---- | ||
| + | [[ECE662:BoutinSpring08_Old_Kiwi|Back to ECE662, Spring 2008, Prof. Boutin]] | ||
| + | [[Category:Lecture Notes]] | ||
Latest revision as of 08:40, 17 January 2013
Lecture 18, ECE662: Decision Theory
Lecture notes for ECE662 Spring 2008, Prof. Boutin.
Other lectures: 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26, 27, 28,
Nearest Neighbors Classification Rule (Alternative Approach)
- Find invariant coordinates
$ \varphi : \Re ^k \rightarrow \Re ^n $ such that $ \varphi (x) = \varphi (\bar x) $ for all $ x, \bar x $ which are related by a rotation & translation Do NOT trivialize!
Example: $ \varphi (x) =0 $ gives us a trivial invariant coordinate. But, you lose information about separation, since everything is mapped to zero.
Want $ \varphi (x) = \varphi (\bar x) $ $ \Leftrightarrow x, \bar x $ are related by a rotation and translation
Example: $ p=(p_1,p_2,\cdots, p_N) \in \Re ^{3 \times N} $ $ \varphi $ maps representation position of tags on body onto $ (d_{12},d_{13},d_{14},\cdots , d_{N-1, N} ) $ where $ d_{ij} $= Euclidean distance between $ p_i $ and $ p_j $
In the above example, we can reconstruct up to a rotation and translation.
WARNING: Euclidean distance in the invariant coordinate space has nothing to do with Euclidean distance or Procrustes distance in initial feature space.
Nearest Neighbor in $ \Re ^2 $ yields tessellation (tiling of floor with 2D shapes such that 1) no holes and 2) cover all of $ \Re ^2 $ ). The tessellations separate sample space into regions. Shape of cells depends on metric chosen. See Figure 1.
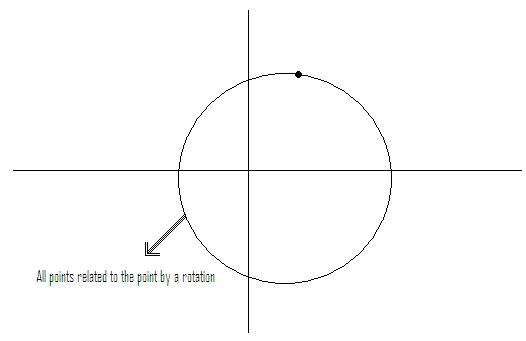
Example: if feature vectors are such that vectors related by a rotation belong to same class
$ \rightarrow $ metric should be chosen so that tiles are rotationally symmetric. See Figure 2.
Instead of working with (x,y) rotationally invariant, work with $ z=\sqrt{x^2 + y^2} $ (distance from origin)
How good is Nearest Neighbor rule?
- Training error is zero: does not measure the "goodness" of a rule
- Test error: want it to be equal to Bayes error rate, because this yields the minimum error
Nearest Neighbor error rate
Recall: Probability of error (error rate) on test data is $ P(e)=\int p(e \mid \vec{x}) p(\vec{x}) d\vec{x} $
Let $ P_d(e) $ be the error rate when d training samples are used.
Let $ P=\lim_{d \rightarrow \infty } P_{d}(e) $
Claim: limit error rate $ P=\int (1-\sum _{i=1} ^{c}p^2 (\omega _i \mid \vec{x}))p(x)dx $
Proof of claim: Given observation $ \vec{x} $, denode by $ \vec{x'}_d $ the nearest neighbor of $ \vec{x} $ among $ \{\vec{x}_1,\vec{x}_2, \cdots , \vec{x}_d \} $
$ P_d(e \mid \vec{x})=\int p_d(e \mid \vec{x}, \vec{x'}_d)p_d(\vec{x'}_d \mid \vec{x})p(x)dx $ but $ \lim _{d \rightarrow \infty } p_d (\vec{x'}_d \mid \vec{x})=\delta ({\vec{x' _d }-\vec{x}}) $
because probability that sample falls into region R centered at $ \vec{x} $ is $ P_{R}=\int _R p(\vec{x' _d})d \vec{x'}_d $.
So, if $ p(\vec{x}) \neq 0 $ (true almost everywhere), then probability that all samples fall outside R is $ \lim _{d \rightarrow \infty} {(1-P_{R})}^d =0 $
So, $ \lim _{d \rightarrow \infty} \vec{x'}_d = \vec{x} $ and $ p_d (\vec{x'}_d \mid \vec{x})=\delta (\vec{x'}_d -\vec{x})+\epsilon _d (\vec{x}) $ where $ \lim _{d \rightarrow \infty } \epsilon _d (x)=0 $
Now $ p_d (e \mid \vec{x}, \vec{x'}_d) $ = ?
Let $ \theta , \theta _1 , \theta _2 , \cdots, \theta _{d} $ be the class of $ x , x_1 , x_2 , \cdots , x_d $, respectively.
Using nearest neighbor rule, error if $ \theta \neq $class of $ \vec{x'}_d $$ =: \theta ' _d $
$ \Rightarrow p_d(e \mid \vec{x},\vec{x'}_d)=1-\sum_{i=1} ^ c p(\theta = \omega _i , \theta ' _d = \omega _i \mid \vec{x}, \vec{x'}_d ) $
$ =1- \sum _{i=1} ^c p(\omega _i \mid \vec{x}) p(\omega _i \mid \vec{x'} _d) $
Recall $ p_d (e \mid \vec{x}, \vec{x'}_d)p_d (\vec{x'}_d \mid \vec{x})d \vec{x'}_d $
You get: $ p_d (e \mid \vec{x})=(1-\sum _{i=1} ^c p(\omega _i \mid x) p(\omega _i \mid x) ) $ + {something that goes to zero as d goes to $ \infty $}
$ \lim _{d \rightarrow \infty} p_d (e \mid \vec{x})=(1- \sum _{i=1} ^c {p(\omega _i \mid x)}^2) $