(→Graph Theory Clustering) |
(→Graph Theory Clustering) |
||
| Line 80: | Line 80: | ||
* A ''minimum spanning tree'' (MST) of a graph G is tree having minimal weight among all spanning trees of <math>G</math>. | * A ''minimum spanning tree'' (MST) of a graph G is tree having minimal weight among all spanning trees of <math>G</math>. | ||
| + | |||
| + | (There are more terminologies for graph theory [http://en.wikipedia.org/wiki/Glossary_of_graph_theory here] ) | ||
Illustartion [[Image:Spark1_OldKiwi.jpg]] | Illustartion [[Image:Spark1_OldKiwi.jpg]] | ||
Revision as of 01:45, 10 April 2008
Clustering Method, given the pairwise distances
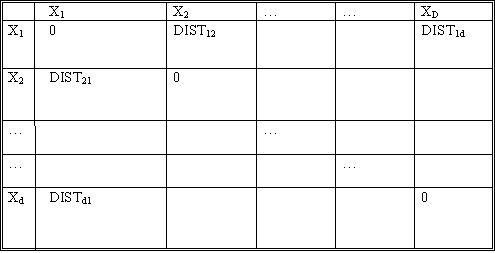
Let d be the number of objects. Let $ DIST_{ij} $ denote the distance between objects $ X_i $ and $ X_j $. The notion of distance here is not clear unless the application itself is considered. For example, when dealing with some psychological studies data, we may have to consult the psychologist himself as to what is the "distance" between given two concepts. These distances form the input to our clustering algorithm.
The constraint on these distances is that they be
- symmetric between any two objects ($ DIST_{ij}=DIST_{ji} $)
- always positive ($ DIST_{ij}>=0 $)
- zero for distance of an object from itself ($ DIST_{ii}=0 $)
- follow $ \bigtriangleup $ inequality
Idea:
If $ DIST_{ij} $ small => $ X_i $, $ X_j $ in same cluster.
If $ DIST_{ij} $ large => $ X_i $, $ X_j $ in different clusters.
- How to define small or large? One option is to fix a threshold $ t_0 $.
such that
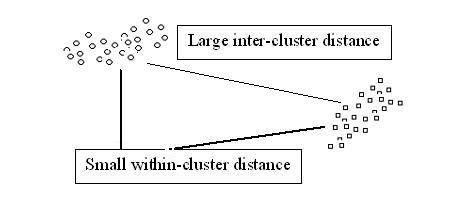
$ t_0 $ < "typical" distance between clusters, and
$ t_0 $ > "typical" distance within clusters.
Consider the following situation of objects distribution. This is a very conducive situation, and almost any clustering method will work well.
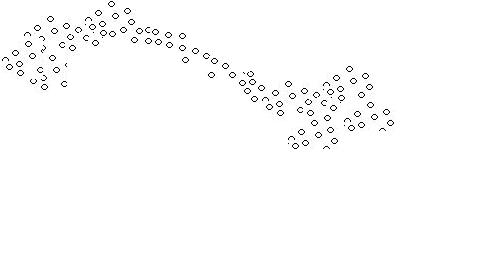
A problem arises when thew following situation arises:
Although we see two separate clusters with a thin distribution of objects in between, the algorithm mentioned above identifies only one cluster.
Graph Theory Clustering
dataset $ \{x_1, x_2, \dots , x_d\} $ no feature vector given.
given $ dist(x_i , x_j) $
Construct a graph:
- node represents the objects.
- edges are relations between objects.
- edge weights represents distances.
Definitions:
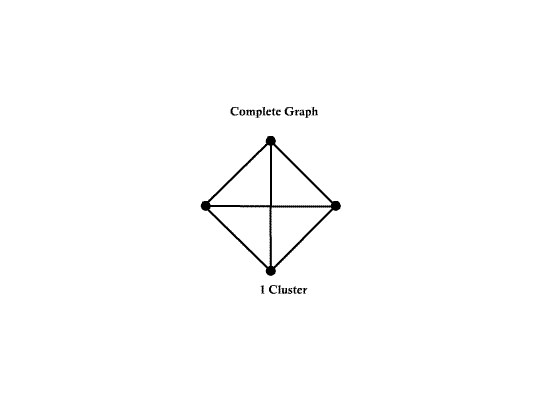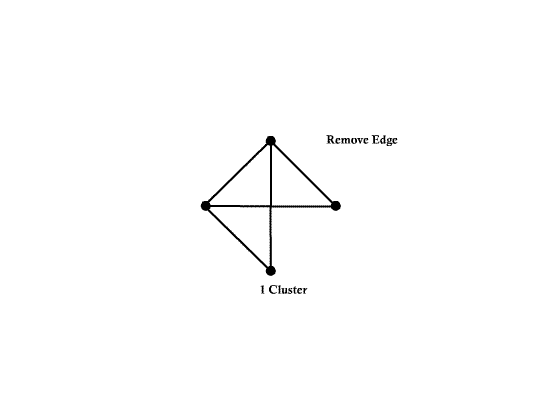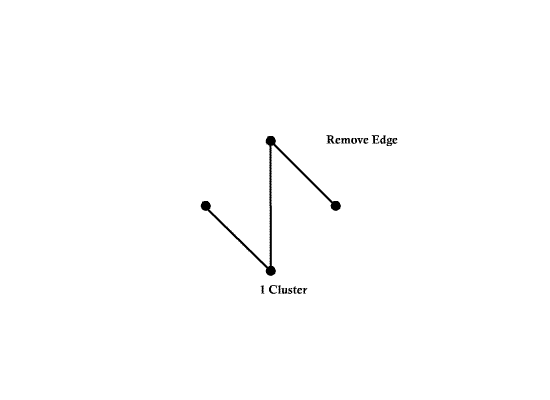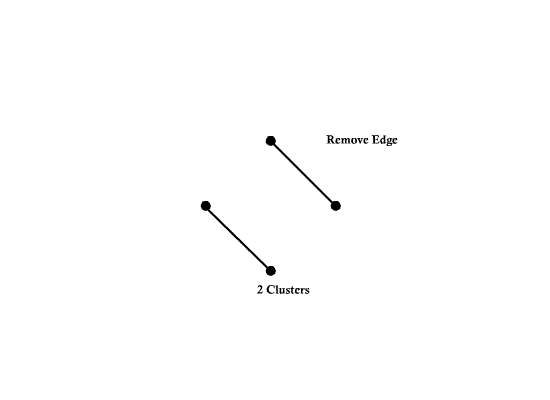
- A complete graph is a graph with $ d(d-1)/2 $ edges.
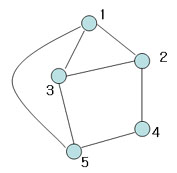
Example:
Number of nodes d = 4 Number of edges e = 6
- A subgraph $ G' $ of a graph $ G=(V,E,f) $ is a graph $ (V',E',f') $ such that $ V'\subset V $ $ E'\subset E $ $ f'\subset f $ restricted to $ E' $
- A path in a graph between $ V_i,V_k \subset V_k $ is an alternating sequence of vertices and edges containing no repeated edges and no repeated vertices and for which $ e_i $ is incident to $ V_i $ and $ V_{i+1} $, for each $ i=1,2,\dots,k-1 $. ($ V_1 e_1 V_2 e_2 V_3 \dots V_{k-1} e_{k-1} V_k $)
- A graph is "connected" if a path exists between any two vertices in the graph
- A component is a maximal connected graph. (i.e. includes as many nodes as possible)
- A maximal complete subgraph of a graph $ G $ is a complete subgraph of $ G $ that is not a proper subgraph of any other complete subgraph of $ G $.
- A cycle is a path of non-trivial length $ k $ that comes back to the node where it started
- A tree is a connected graph with no cycles. The weight of a tree is the sum of all edge weights in the tree.
- A spanning tree is a tree containing all vertices of a graph.
- A minimum spanning tree (MST) of a graph G is tree having minimal weight among all spanning trees of $ G $.
(There are more terminologies for graph theory here )
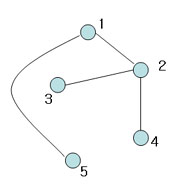 This is spanning tree (MST with weight 4)
This is spanning tree (MST with weight 4)
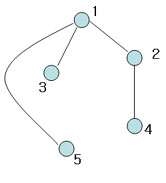 Another spanning tree (Also, MST with weight 4)
Another spanning tree (Also, MST with weight 4)
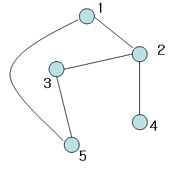 Not spannning tree (Include 'cycle')
Not spannning tree (Include 'cycle')
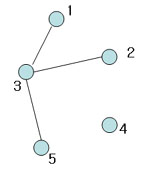 Not spanning tree (No edge for 4)
Not spanning tree (No edge for 4)
Graphical Clustering Methods
- "Similarity Graph Methods"
Choose distance threshold $ t_0 $
If $ dis(X_i,X_j)<t_0 $ draw an ege between $ X_i $ and $ X_j $
Example) $ t_0= 1.3 $
Fig 3-1
Can define clusters as the connected component of the similarity graph
=> Same result as "Single linkage algorithms"
$ X_i \sim X_j $ if there is chain
$ X_i \sim X_{i_1} \sim X_{i_2} \sim \cdots X_{i_k} \sim X_{j} $ complete
Can also define clusters as the maximal subgraphs of similarity graph
Fig 3-2
=>this yeilds more compact, less enlongated clusters
The approach above is not good for, say,
Fig 3-3